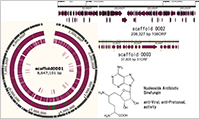
Enlarge Image Draft genome sequencing is completed and assembled on one major scaffold0001 which is linear genome containing 8,647,101bp, with telomere sequences at both of the ends. Two additional plasmids scaffold 0002 and 0003 were also elucidated by the assembled contigs.
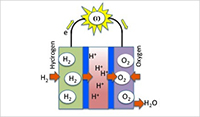
Enlarge Image A biofuel cell is a battery system in which a pair of enzymes works together to generate power by oxidizing hydrogen gas (H2) on one side and reducing oxygen gas (O2) on the other side. The anode inevitably requires hydrogenase while the cathode reserves choices of enzymes including formate dehydrogenase.
Enlarge Image
Genome sequencing anti-viral antibiotics-producing Streptomyces incarnatus, led to the discovery of Se-containing formate dehydrogenase gene for biofuel cell
Despite the general understanding that antibiotics cannot be prescribed to cure the common flu, there are lines of antibiotics that are capable of killing viruses, protozoa, fungi, and even cancer cell lines. Notably, these antibiotics-referred to as “nucleoside antibiotics”- are not utilized in our society due to the extremely poor production yield by usual fermentation processes.
Here, in order to exploit the genetic codes of nucleoside antibiotics-producing Streptomycs incarnatus NRRL8089, Takashi Tamura and colleagues at Tokyo University have elucidated the 8,897,465-bp genome sequence that were assembled in computer simulations, and putative genes were annotated by homology search, and the sequence opened to public in the international database NCBI.
The Gram-positive bacterium is now being investigated with the aim of dramatically improving production through genome engineering via de-regulation of gene transcription by an RNA polymerase (rpoB mutation) and protein quality-control system consisting of groES/groEL chaperoning system, which ultimately improves the thermal stability of the biosynthetic enzymes and thereby endures the long-lasting production of these precious antibiotics and metabolites.
Intriguingly, a serendipitous discovery from these genome sequencing was an open reading frame for selenocysteine-containing formate dehydrogenase-O (fdh-O gene) together with the set of the selenoprotein biosynthetic gene cluster selABCD in close vicinity on the bacterial genome.
Importantly, the selenoprtein Fdh-O has potential applications for the development of bio-fuel cells for generating electricity from hydrogen without burning fossil fuels.
Genome sequencing may offer clues into the efficient and convenient mass production of nucloside antibiotics for large-scale cathode enzymes for the development of biofuel cells.
Reference1:
・Authors: Kenshiro Oshima, Masahira Hattori, Hitomi Shimizu, Koji Fukuda, Michiko Nemoto, Kenji Inagaki, Takashi Tamura
・Title of original paper: Draft genome sequence of Streptomyces incarnatus NRRL8089,which produces the nucleoside antibiotic, Sinefungin
・Journal, volume, pages and year: Genome Announcements 3, e715 (2015).
・Digital Object Identifier (DOI): 10.1128/genomeA.00715-15
・Journal website: http://dx.doi.org/10.1128/genomeA.00715-15
・Journal, volume, pages and year: Genome Announcements 3, e715 (2015).
http://ousar.lib.okayama-u.ac.jp/metadata/53727
・Affiliations: Graduate School of Life and Environmental Sciences, Okayama University, Okayama, Japan.